Notice
MMODA multi-messenger online data analysis platform in the frame of the EuroScienceGateway project
- document 1 document 2 document 3
- niveau 1 niveau 2 niveau 3
Descriptif
The platform makes use of cloud-based data management solutions and virtualization technologies. It is intended to meet the challenges of efficient sharing and reuse of data, long-term preservation of data and data analysis workflows and the reproducibility of results. Some analysis modules (like the one for the INTEGRAL data analysis) developed as separate plugins. In addition to these “fixed” plugins, the analysis workflows in the form of Jupyter notebooks, annotated by the terms from our platform-specific ontology, can be automatically deployed as a service with web-GUI and REST API.
The MMODA platform is based on a federated computing environment. Each instance is based on a layer of physical or virtual machines running microservices in the containerized form. For the deployment, we use Kubernetes container orchestrator, which is automatically deployed by the Rancher Kubernetes Engine (RKE2) solution via the TerraForm module on top of OpenStack IaaS cloud of the French academic cloud federation, FG-cloud. Helm charts are provided to deploy all MMODA services (web interfaces and API, task manager, adapters corresponding to scientific analyses, etc.). We use the GitOps practices to deploy the platform software stack.
Many approaches to data management and workflow publishing are rather close between communities, but realisations are completely separate. As an example, the BioInformatics community has somewhat widely adapted Galaxy platform, enabling open data analysis services with distributed compute. We joined (along with particle physics, material science, climate science) an EuroScienceGateway EOSC project in order to bring our tools and workflows into Galaxy platform, developing synergy between MMODA and Galaxy. We explore peculiarities of the astroparticle-oriented workflow publishing on the WorkflowHub platform. This project, as follows, allows minimizing redundancy and enable cross-domain knowledge transfer.
Intervention / Responsable scientifique
Thème
Documentation
Dans la même collection
-
Table ronde et discussions : infrastructures de calcul et ateliers de la donnée de recherche Data G…
CastexStéphanieRenardArnaudAlbaretLucieRenonNicolasDufayardJean-FrançoisPARTIE 1 : Introduction et présentation des intervenants
-
Table ronde et discussions : infrastructures de calcul et ateliers de la donnée de recherche Data G…
CastexStéphanieRenardArnaudAlbaretLucieRenonNicolasDufayardJean-FrançoisPARTIE 6 : L'accompagnement autour de la gestion des données
-
Table ronde et discussions : infrastructures de calcul et ateliers de la donnée de recherche Data G…
CastexStéphanieRenardArnaudAlbaretLucieRenonNicolasDufayardJean-FrançoisPARTIE 3 : les liens entre les structures
-
Table ronde et discussions : infrastructures de calcul et ateliers de la donnée de recherche Data G…
CastexStéphanieRenardArnaudAlbaretLucieRenonNicolasDufayardJean-FrançoisPARTIE 5 : Des interactions croisées, à propos des compétences
-
Table ronde et discussions : infrastructures de calcul et ateliers de la donnée de recherche Data G…
CastexStéphanieRenardArnaudAlbaretLucieRenonNicolasDufayardJean-FrançoisPARTIE 2 : Les ateliers de la données
-
Table ronde et discussions : infrastructures de calcul et ateliers de la donnée de recherche Data G…
CastexStéphanieRenardArnaudAlbaretLucieRenonNicolasDufayardJean-FrançoisPARTIE 7 : Conclusion
-
Table ronde et discussions : infrastructures de calcul et ateliers de la donnée de recherche Data G…
CastexStéphanieRenardArnaudAlbaretLucieRenonNicolasDufayardJean-FrançoisPARTIE 4 : Des interactions croisées, liens entre le calcul et les données.
-
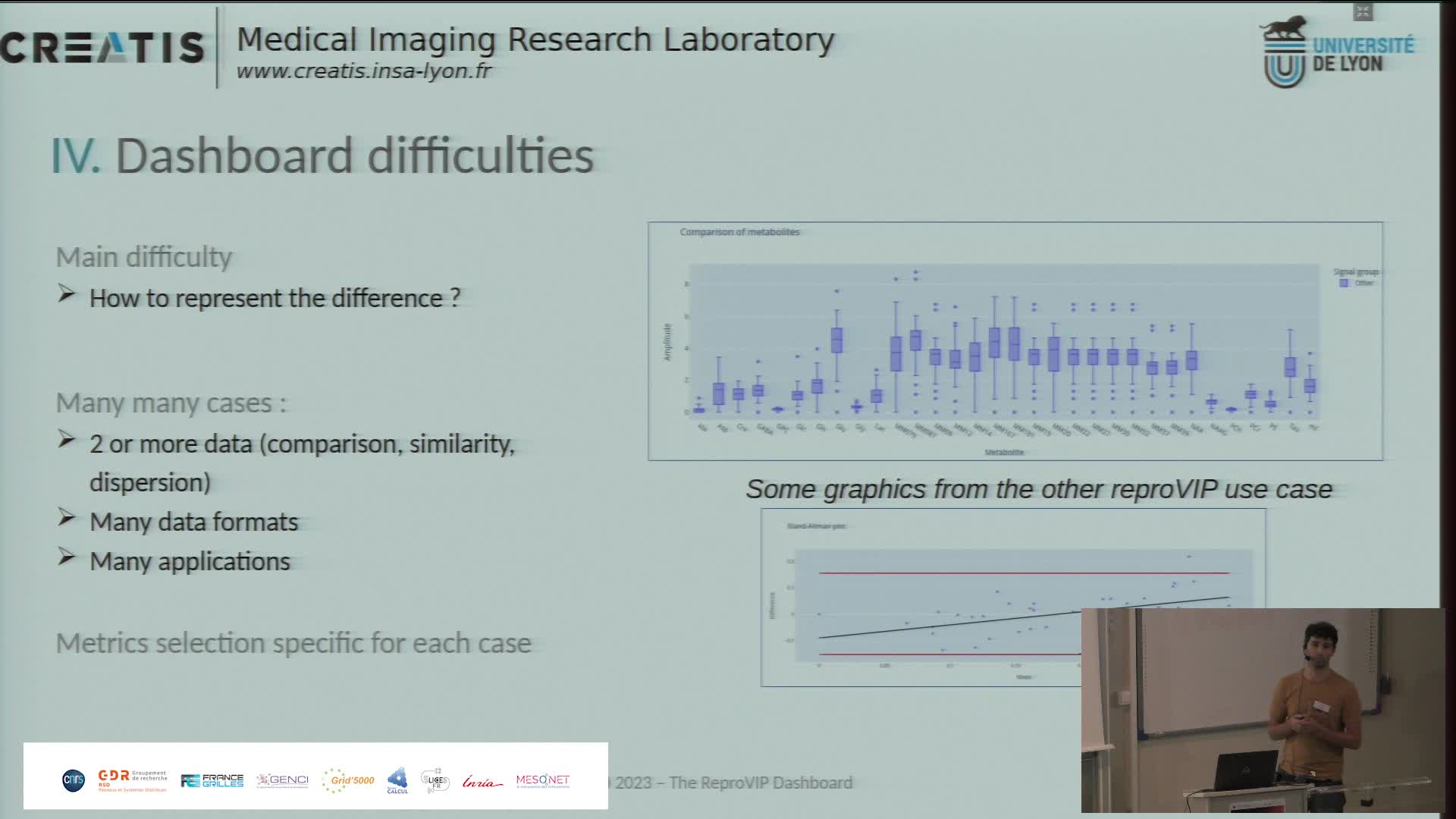
Présentation du dashboard ReproVIP pour visualiser la reproductibilité dans l'imagerie médicale
BonnetAxelLa plateforme d'imagerie virtuelle VIP [1] (https://vip.creatis.insa-lyon.fr) est un portail web de simulation et d'analyse d'images médicales. Elle existe depuis plus de 10 ans et a évolué pour
-
Parallélisation par l'intermédiaire d'une fenêtre à mémoire partagée (MPI 3.0) : application à un c…
ElyakimePierreJADIM est un code de calcul de mécanique des fluides développé en Fortran 90 à l'Institut de Mécanique des Fluides de Toulouse (IMFT).
-
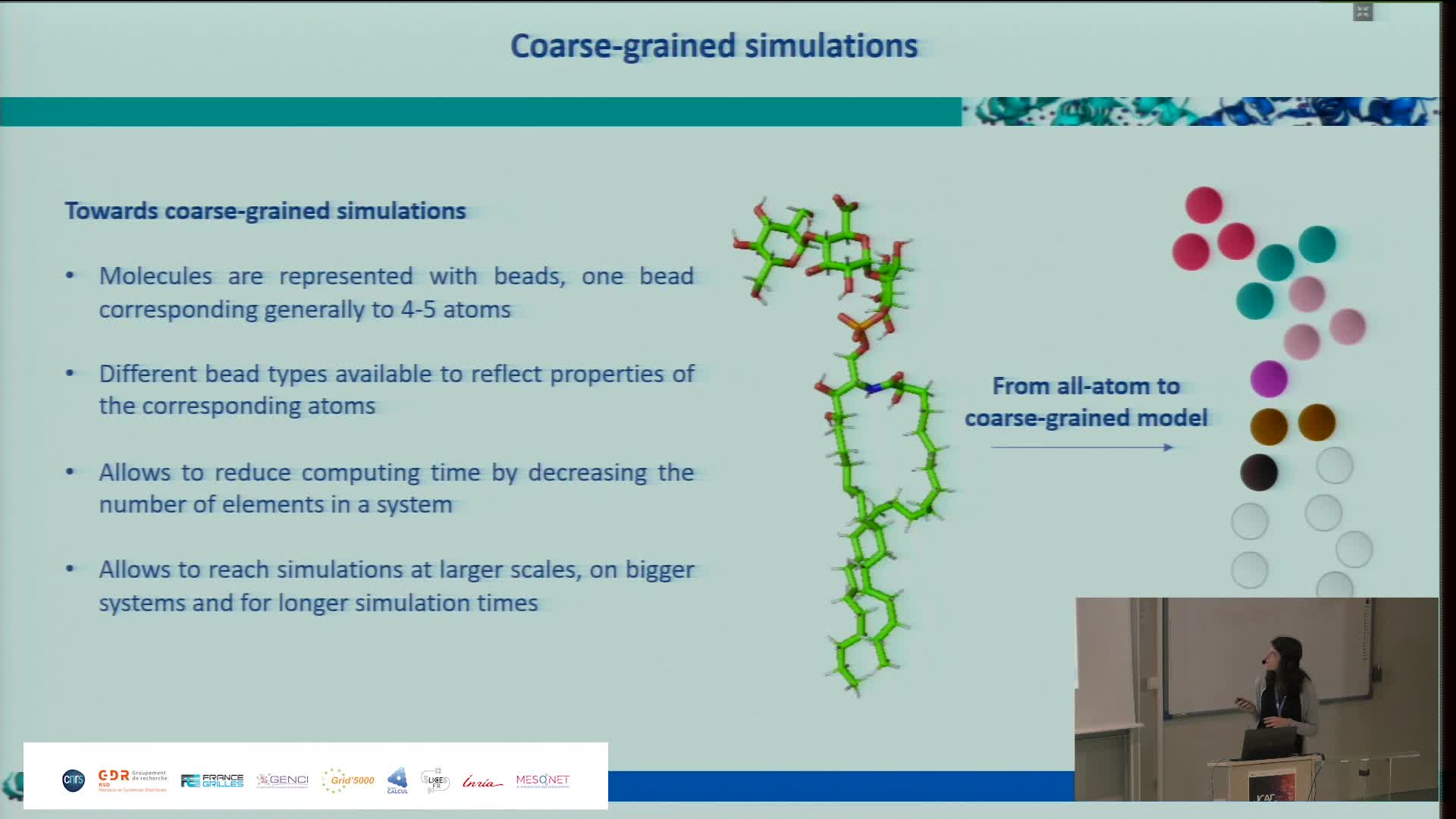
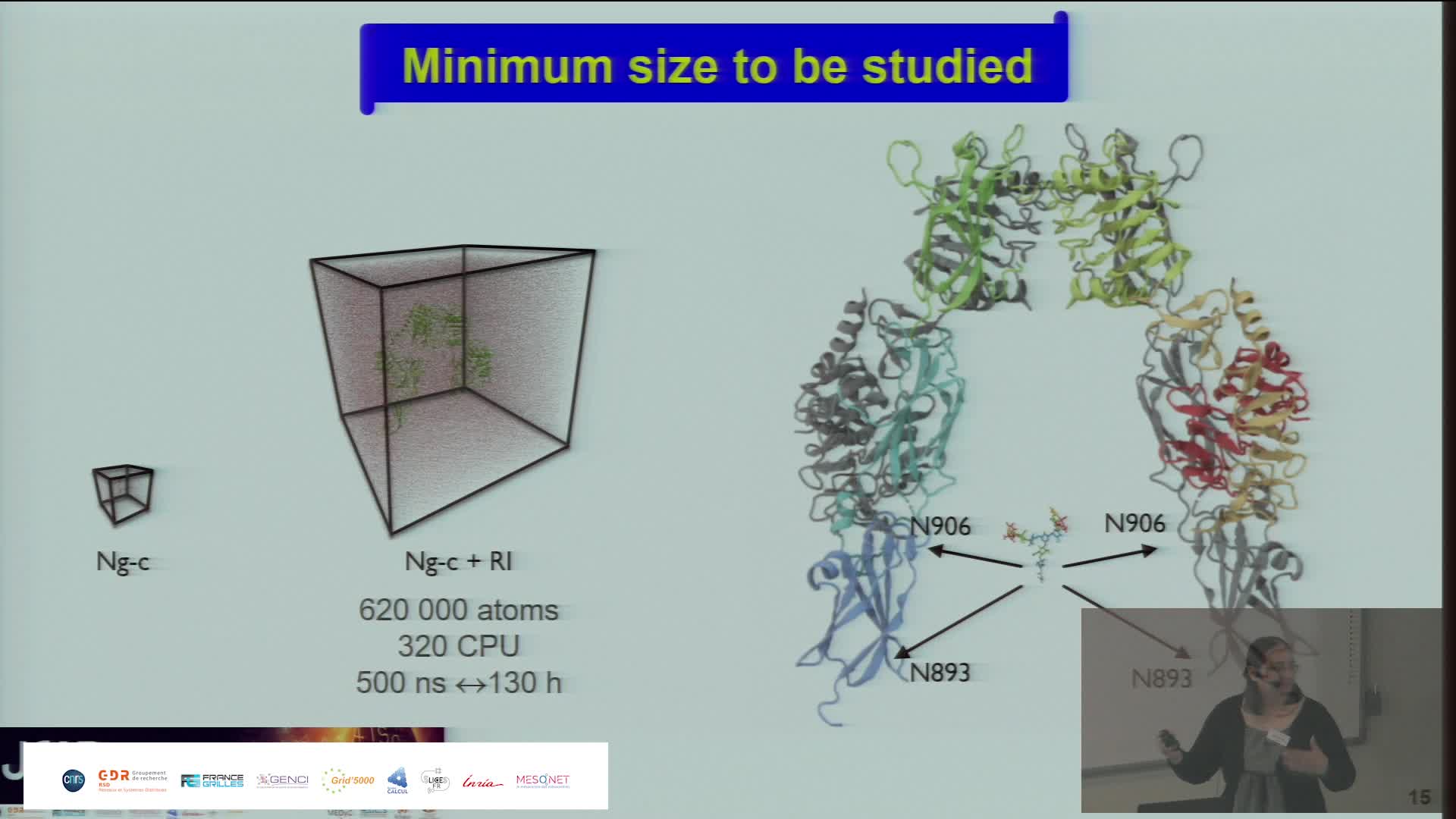
Simulating the Extra Cellular Matrix - Calculations and data from atom to animal
BaudStéphanieThe extracellular matrix (ECM) is a three-dimensional network of macromolecules that is the architectural support for cells and allows tissue cohesion. This dynamic structure regulates many biological
-
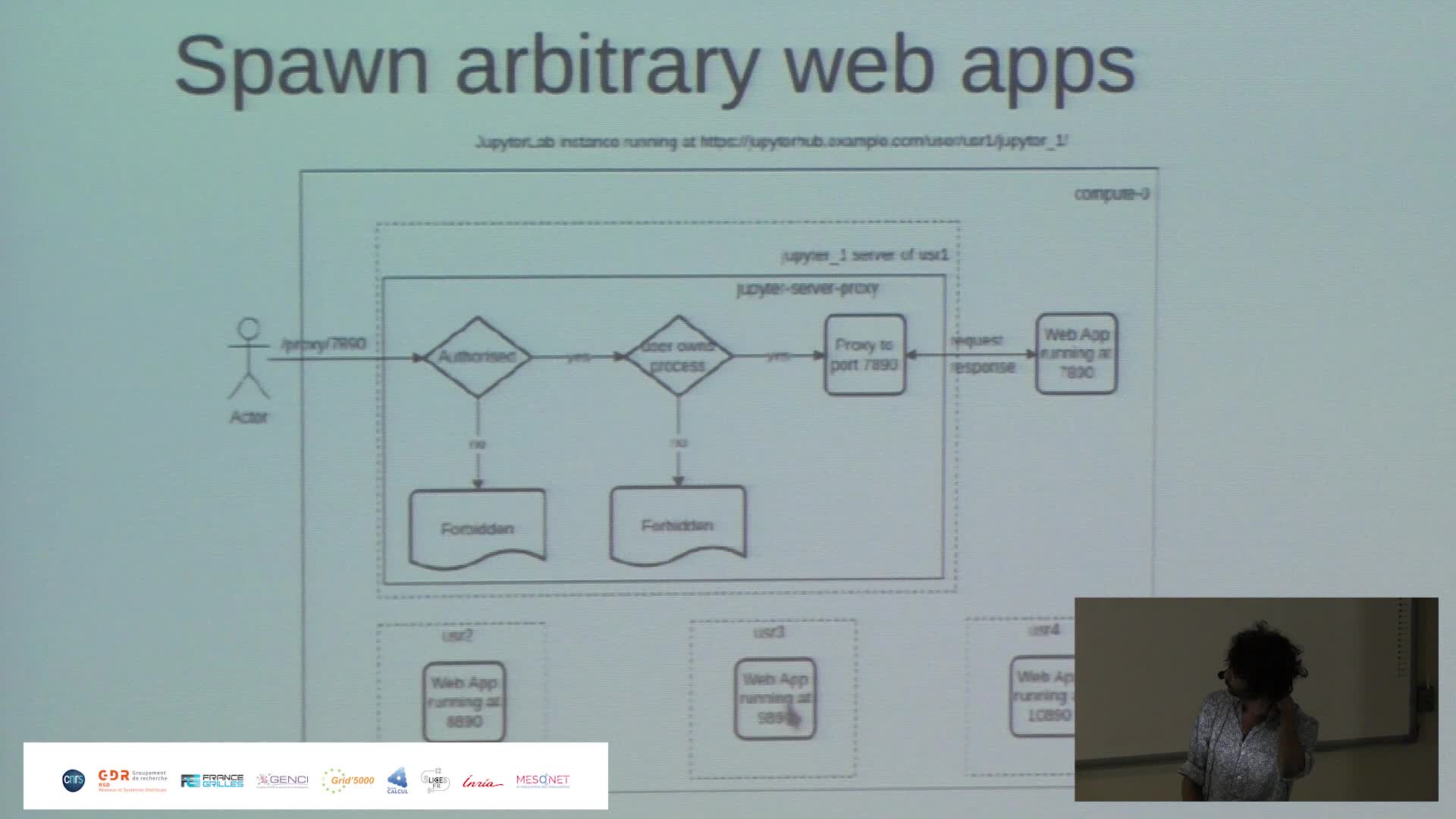
Harnessing the power of Jupyter{Hub,Lab} to make Jean Zay HPC resources more accessible
PaipuriMahendraThis talk revolves around the deployment of JupyterHub on Jean Zay HPC platform and how it is done to meet the demands of RSSI (ZRR constraints) and also to provide a seamless experience to the end
-
Using High Performance Computing to decipher the impact of ageing process on collagens
DepenveillerCamillePlant cells have to face environmental stress, and in this context, being able to decipher the organization of the plant plasma membrane is a key point to better understand this adaptability of plasma